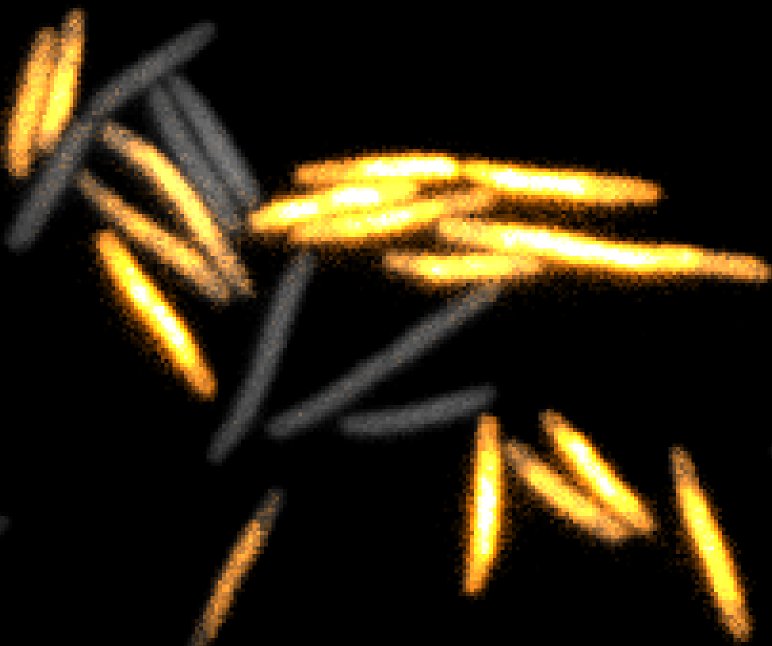DNA repair, Mutation, and Stress Adaptation in Bacteria
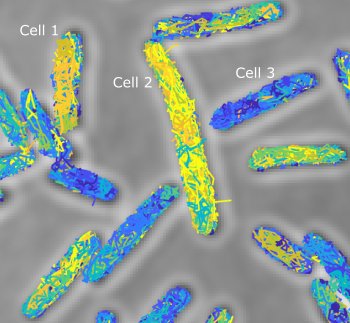
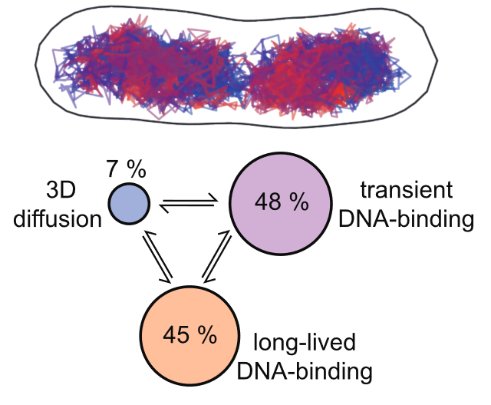
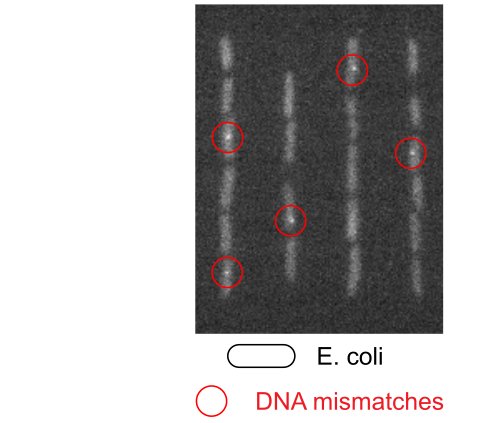
Bacteria have a remarkable capacity to thrive in adverse environments. Their adaptability relies on stress responses that provide temporary protection, for example by repairing cell damage or removing toxic chemicals. Such phenotypic adaptation offers cells a window of opportunity to evolve permanent stress resistance through genetic change. Failures to cure bacterial infections with antibiotics are often due to stress responses that promote bacterial survival as well as the evolution of drug resistance. Our lab seeks to understand how this works at the molecular level using a quantitative interdisciplinary approach. We focus on the mechanisms of DNA repair and mutagenesis, which are essential both for stress survival and for genetic change. A key aspect of our research is developing fluorescence microscopy techniques to visualise molecular events in real-time within living cells. We use super-resolution microscopy and single-molecule tracking to record the localization and movement of individual molecules such as DNA repair enzymes or transcription factors. To monitor the cellular responses to stress, we use microfluidic devices for imaging single cells. This allows us to decipher how molecular events inside cells determine long-term cell fates. Curiously, single-cell analysis revealed that bacterial phenotypes are variable even in a constant environment, a phenomenon that may be linked to stress survival. We discovered that mutation rates are also variable due to fluctuations in the expression of DNA repair proteins. These findings open fundamental questions about the mechanisms and regulation of mutagenesis, which we are now addressing using a range of novel microscopy and genetic approaches.
Uphoff Lab | Department of Biochemistry | University of Oxford

News
Chaotic behaviour
December 2023
Single bacteria exhibit chaotic oscillations in gene expression in response to oxidative stress. Our new study in Current Biology demonstrates that chaos exists in biological systems and may play a role in stress adaptation of cell populations.
A burst of mutations
November 2022
Cell stress triggers phenotypic and genetic changes. Our paper in EMBO Reports shows how stress responses modulate mutation rate dynamics: Adaptation delay causes a burst of mutations in bacteria responding to oxidative stress.
Phenotypic heterogeneity driven by cell-cell interactions
September 2022
Divya's paper in Cell Reports shows that Phenotypic heterogeneity in the bacterial oxidative stress response is driven by cell-cell interactions
Regulation of the oxidative stress response
November 2024
How does one transcription factor control over 20 genes to protect bacterial populations from oxidative stress? Check our new paper in Cell Systems for some answers: The master regulator OxyR orchestrates bacterial oxidative stress response genes in space and time
DNA interstrand crosslink repair visualised in cells
June 2023
Our lab contributed to a paper by Martin Cohn's group on the regulation of DNA interstrand crosslink repair, now published in Cell Reports: Phosphorylation by ATR triggers FANCD2 chromatin loading and activates the Fanconi anemia pathway
Book chapter
August 2022
Chloe Cassaro's chapter in the series Methods in Molecular Biology includes a detailed point-by-point protocol for Super-Resolution Microscopy and Tracking of DNA-Binding Proteins in Bacterial Cells
Imaging outer membrane biogenesis
February 2025
Our team contributed to research on how lipopolysaccharides and outer membrane proteins are distributed in growing bacteria. This work was led by the Kleanthous lab and is now published in PNAS!
Improving single-molecule tracking methodology
June 2024
Using live-cell single-molecule tracking for your experiments, but not sure which fluorescent label to use? Our new study might help!
Anju Kudhail and Elliott Haywood
September 2024
Anju and Elliott have joined our team as DPhil students in the DTP programme.