MICRON BIOIMAGING FACILITY
Access to advanced imaging technology
Including bespoke development systems
With expert guidance and support for your project
Access to advanced imaging technology
Including bespoke development systems
With expert guidance and support for your project
Accesso effettuato come:
filler@godaddy.com

We help researchers apply cutting edge fluorescence microscopy technology to their research questions. See our range of microscopes here.
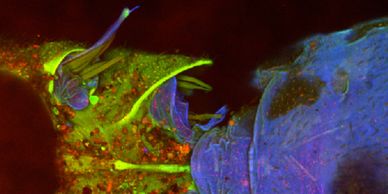
Find out how to access our cutting edge imaging technology including our super-resolution and development systems.

Wide ranging high profile publications from addressing key biological questions to development of new instruments and approaches.

Find out our latest events and news happening in the facility.

Access to all our fluorescence microscopy and image analysis educational tools.
3D rendered movie of a kidney section captured using a Leica confocal microscope. Alexa Fluor® 488 wheat germ agglutinin is a fluorescent lectin that labels the glomeruli and convoluted tubules (cyan). Actin located in glomeruli and the brush border are stained with Alexa Fluor® 568 phalloidin (magenta), nuclei are stained with the DNA stain DAPI (grey).
3D rendered movie of the same sample on the left captured using a Leica confocal microscope. The data has been deconvolved using the Lightning algorithm to remove noise. Data was captured using a 63x NA1.1 oil objective.

The primary source of funding for our cutting-edge imaging research.
Principal Applicant: Ilan Davis
Co-applicants: Jordan Raff, Martin Booth, Yvonne Jones, Christian Eggeling, David Stuart, Kay Grunewald, Neil Brockdorff.

Funded by the Wellcome Institutional Strategic Support Fund and the John Fell Fund, these two innovative microscopy development projects, the 4Pi SMS & Microscopi, are aimed at very different areas of research.
Our microscopes are housed at the Department of Biochemistry, in Phase 1 of the New Biochemistry building.
South Parks Rd, Oxford, OX1 3QU, UK